I believe that humans represent the ideal model organism for the development of innovative therapies to improve human health. Experiments of nature (e.g., human genetics) and longitudinal observations in patients with disease can differentiate between cause and consequence, and therefore can overcome fundamental challenges of drug development (e.g., target identification, biomarkers of drug efficacy). Using my Twitter account (@rplenge), this blog (www.plengegen.com/blog), and other forms of social media, I provide compelling examples that illustrate key concepts of “humans as the ideal model organism” (#himo) for drug development.
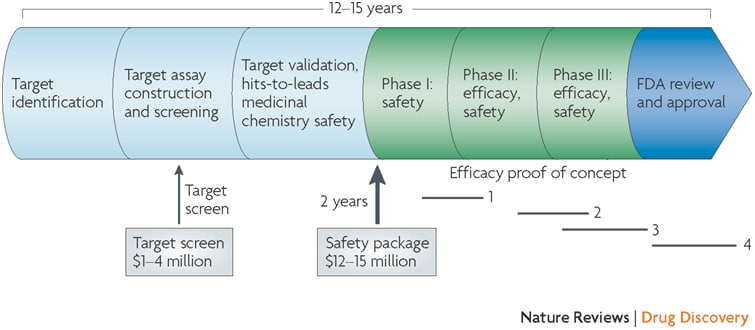
Why do drugs fail (#whydrugsfail)? This simple question is at the center of problems facing the pharmaceutical industry. In short, drugs fail in early development because of unresolved safety signals or lack of biomarkers for target engagement, and drugs fail in late development because of lack of efficacy or excess toxicity. This leads to a costly system for bringing new drugs to market – not because of the successes, but because >95% of drug programs ultimately fail. Without improvements in rates of success in drug development, the sustainability of the pharmaceutical industry as we know it is in trouble (see here). Not surprisingly, much has been written about this topic, including analyses of development strategies (Forbes blog, Drug Baron), company pipelines (Nature Reviews Drug Discovery manuscript from AstraZeneca) and FDA approvals (here and here).
So why do we need yet another blog or twitter account that highlights the challenges of the pharmaceutical industry? My answer is that I will take a unique angle to potential solutions, with a focus on experiments and observations derived from the ideal model organism – humans. These examples will be filtered through my experience in clinical medicine (MD degree [CWRU], academic training in internal medicine [UCSF], rheumatology [Brigham and Women’s Hospital], and clinical bioinformatics [Partners HealthCare]); genetics and genomics (PhD in genetics [CWRU], post-doctoral research [Broad Institute] and independent faculty / principle investigator [Harvard Medical School]; and the pharmaceutical industry (Vice President and head of Genetics & Pharmacogenomics [Merck]).
Here are examples of subjects that I will cover in my social media posts, which I am sure will evolve over time:
(1) Human genetics (#genetics)
In late development, most drugs fail due to lack of efficacy. This occurs despite an abundance of pre-clinical data implicating a target in disease. That is, drugs fail in late development because we pick the wrong targets from the outset. As I have written before (see here, here), and others have as well (see here, here), human genetics represents a powerful way to pick targets and mechanisms of action at the earliest stage of drug discovery.
One way to think about human genetics in drug discovery is to use the same framework that defines a successful drug: dose-response curves. In a Nature Review Drug Discovery article that I wrote with David Altshuler (Harvard Medical School & Broad Institute) and Ed Scolnick (former President of Merck Research Laboratories), we developed the argument of “genotype-phenotype dose response curves” for single genes associated with human traits (see original manuscript here). The manuscript provides criteria for nominating drug targets based on human genetics. In a previous blog post (see here), I applied these criteria to PCSK9, which represents one of the best examples of genetics to drive drug discovery.
Going forward, I will use this blog and Twitter to highlight examples of human genetics in action. I expect many examples to be new discoveries, as recently shown for APOC3 and myocardial infarction (see here, here, here). I also expect many posts to about functional consequences of human mutations, including molecular studies on gene/mutation mechanism of action, emerging technologies such as CRISPR-Cas9, and methodological approaches such as Mendelian randomization (see here) or rare variant discoveries (see here, here, and Daniel MacArthur lab link here). Other posts will relate to technological innovations in human genetics such as clinical sequencing (here).
[If you want to see a slide deck and blog that I put together for a course on genetics in immunology at Harvard Medical School, see here; if you want a slide deck that I prepared for the New York Academy of Sciences on how genetics changes the approach from target identification to the clinic, see here.]
(2) Pharmacogenomics (#pgx)
Pharmacogenomics is a broad field. I define it as the use of genetic and genomic biomarkers to understand how drugs work: biomarkers to monitor drug-target engagement; biomarkers to define novel mechanisms of action; biomarkers to guide indication selection, and biomarkers to predict efficacy and toxicity in clinical trials. This definition goes beyond straightforward pharmacogenetics (see nice review article on clinical pharmacogenetics by David Goldstein here) and also beyond the ambiguous field of “personalized medicine” (see my previous blog here).
In future blog and Twitter posts, I will showcase examples of pharmacogenomics – with a perspective on successes and failures in drug development. Some posts will relate to genetic predictors of drug efficacy and toxicity, as in the evolving story of genetic predictors of warfarin metabolism in clinical care (see 2013 NEJM articles here, here). Other posts will relate to “big data” in precision medicine, although this topic has plenty of coverage elsewhere, too. I expect to dive deeply into the science as well, including new areas such as polygenic predictors of treatment response (see here) or crowdsourcing approaches to analysis of complex datasets (see here for an example of a project I launched with Stephen Friend at Sage Bionetworks).
One area I will follow closely relates to pharmacogenomics in immuno-therapy, which is in the midst of revolutionizing treatment of patients with cancer (see Matthew Herper’s Forbes blog here). For example, new genomic technologies such as T cell or B cell repertoire sequencing offer an innovative strategy to profile the human immune system. Coupled with other technologies such mRNA expression profiling, mass-spectrometry-based proteomic profiling and next-generation sequencing, detailed molecular observations in humans are now possible.
(3) Next-generation clinical registries (#nextgenreg)
Just as the current model of the pharmaceutical industry is being challenged, the current model of clinical registries is entering a disruptive phase. The cost of most registries, together with the need for extremely large patient cohorts, is changing the way patients are enrolled in discovery research studies. I believe this topic is highly relevant to the theme of “humans as the ideal model organism” (#himo) for drug discovery, as a critical first step in any human discovery study is access to appropriate patient samples.
In future blog and Twitter posts, I will opine on the role of electronic medical records in discovery research (which I experienced during may days with the NIH-funded program i2b2, see publications here and i2b2 website here; NPR commentary here); open-source solutions for genome-sequencing data (Global Alliance for Genomics & Health here and here, Personal Genome Project here); patient-oriented registries (such as the arthritis internet registry, which I co-founded in 2010, or direct-to-consumer companies such as 23&Me); and innovated strategies to link genetic data to patient outcome data, especially for the purpose of cataloguing human knock-outs (see here), conducting phenome-wide association studies (PheWAS, see manuscript here and David Grainger’s commentary on FierceBiotech here), defining unusual outcomes (e.g., the The Resilience Project), or establishing unique biobank resources (see here, here)
(4) Innovation in complex organizations (#innovation)
The sheer complexity of the problem – humans as the model organism for drug discovery – necessitates complex organizations. As a result, I believe it is crucial to be innovative and creative in a complex organization, whether that is a pharmaceutical company, an academic institution, biotech, non-profit, or another organization. Beyond my current position at Merck (see description of our Genetics & Pharmacogenomics [GpGx] Department here), I have been fortunate to witness first-hand the launch of highly successful organizations (e.g., Broad Institute), serve leadership positions on complex teams in academia (e.g., i2b2, Pharmacogenomics Research Network [PGRN], Human Immunology Center at Brigham and Women’s Hospital), and participate in strategy development for non-profit organizations (e.g., American College of Rheumatology, Arthritis Foundation, Rheumatology Research Foundation). Moreover, I have had the chance to participate in a wide-variety of consortia and collaborations, including the strategy (while at Harvard) and implementation (while at Merck) of the NIH-industry Accelerating Medicines Partnership (AMP, see here). Through these experiences, I have come to appreciate the importance of an organizational structure on solving important problems such as maximizing discovery research in humans for drug discovery.
In future blog and Twitter posts, I will provide a creative perspective on innovation in complex organizations. For example, what is the relevance of a “fast fail” strategy in drug discovery (see excellent Drug Baron post)? What is the nature of academic collaborations and the future of medicine (see Harvard Medical School Dean Jeff Flier commentary here)? What drug discovery lessons can be learned from other complex organizations such as Pixar (see thoughtful David Shaywitz blog post here), the Panama Canal (see here) or professional sports (see Bruce Booth blog post here and Kyle Serikawa post here)? What can DARPA and Netflix teach us about innovative organizations (see here and here)? How does a leader create a community that is willing and able to innovate (see HBR article on Collective Genius)?
If these or related themes appeal to you, please join me (@rplenge and www.plengegen.com/blog) in an exciting conversation on humans as the model organism of choice in drug discovery! I promise to keep the conversation highly relevant to drug discovery (for example, no posts of my children or contentious political commentary).
I want to acknowledge that I am currently an employee of Merck. The views and opinions presented here and in future blogs do not necessarily reflect those of Merck. Moreover, I will be very careful not to present any material information about the Merck pipeline that is not available in the public domain.
Finally, I want to briefly describe my motivation for these social media posts, which includes psychological forces (see here). First, I want to influence the way drug discovery is done now and in the future – even if it is a very, very small influence! I don’t think that the full power of “humans as the model organism of choice” in drug discovery is fully appreciated. In particular, I don’t think that the application of human genetics to drug discovery has been maximized. Further, there are common misconceptions about the role of genetics in drug discovery that need clarification. The deluge of human genetic data in the near future will make this discussion of even greater importance. Second, I am intrigued by the power of networks, and I want to experience first-hand how social media can be used to advance a cause that I believe in strongly. And third, I think the pharmaceutical industry gets a bad rap on being open with regards to scientific discoveries. In a small way, I can contribute to more transparency in the industry – not by directly revealing trade secrets, but by providing examples from the public that the industry uses to make decisions on drug development.
I hope that my social media presence will give a new voice to the exciting world discovery research in humans as it pertains to drug development at Merck and beyond.