[I am an employee of Celgene. All opinions expressed here are my own.]
A meeting was recently convened to discuss a roadmap for understanding the genetics of common diseases (search Twitter for #cdcoxf18). I presented my vision of a genetics dose-response portal (slides here; link to related 2018 ASHG talk here). The organizers (@RachelGLiao, @markmccarthyoxf, @ceclindgren, Rory Collins [Oxford], Judy Cho [New York], @NancyGenetics, @dalygene, @eric_lander) asked participants to share their vision. I thought I would blog about my mine.
You’ll notice my vision is ambitious. Nonetheless, I believe these objectives are feasible to accomplish within a 3-year (Phase 1) and 7-year (Phase 2) time frame. Phase 1 would start immediately and would guide projects for Phase 2. In reality, many aspects of Phase 1 are already underway today (e.g., GWAS catalogue at EBI; Global Alliance for Genomics and Health [GA4GH] data sharing methods). Phase 2 consists of two parts: federation of global biobanks and experimental validation of variants, genes and pathways. Some components of Phase 2 could start today (e.g., exome sequencing in >100,000 cases selected from existing case-control cohorts and biobanks; human knockout project). As with Phase 1, many components of Phase 2 are already underway (federation of existing biobanks [e.g., UK Biobank with FinnGen], technologies for high-throughput CRISPR mutagenesis and single-cell eQTL analysis).
You’ll also notice my vision is biased towards target-driven drug discovery. However, the resources generated would enable much more than just a dose-response portal for drug discovery (e.g., genetically-validated cellular readouts for phenotypic screens; catalogue of nodes not targeted by existing therapies; hypothesis-generation for novel therapeutic modalities). Perhaps more importantly, the resources would enable much, much more than just drug discovery.
- Phase 1 – All x All Association study (AAAS)
- organize summary statistics for all existing GWAS data (including disease traits and quantitative traits such as mRNA expression, protein levels, peripheral blood counts)
- fine-map signals of association for each trait
- nominate “causal” variant(s) and gene(s) for each trait (using epigenetic data, human cell atlas, coding variants, etc.)
- co-localize all signals of association across all traits (i.e., all x all association study)
- for disease traits co-localized with quantitative traits (e.g., eQTLs, pQTLs), estimate magnitude of clinical effect size relative to quantitative trait effect size
- for all disease traits, nominate pathogenic cell types using epigenetic, gene expression data, etc.
- for all traits, generate polygenic scores (PS) and perform all x all PS association study
- integrate all confirmed rare mutations associated with Mendelian phenotypes
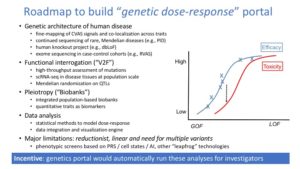
- for up to 1000 “solved genes” with more than one independent associated allele (common or rare, coding or non-coding), estimate genotype-phenotype dose-response curves
- perform Mendelian randomization for disease-associated variants from 1000 “solved genes”
- create searchable database, including visualization tools, for all results, using platforms such as the Open Targets genetics portal
- implement Global Alliance for Genomics and Health (GA4GH) data sharing methods and create data sharing infrastructure for all future GWAS studies
- create modular, open-source, automated pipeline for all components above to incentive future investigators to deposit data
- nominate studies for Phase 2 (e.g., single-cell eQTL studies in pathogenic cell types in >500 individuals, exome sequencing in >100,000 cases from up to 10 selected diseases and/or quantitative traits, high-throughput CRISPR perturbations in relevant cellular systems, human knock-out project)
Rationale for Phase 1: summary statistics are only available for ~30% of GWAS data; poor incentive structure for investigators to make summary statistics and individual level genotype data available; no standardize data sharing format; most large-scale omics data resides in silos; as a consequence, current approach for data integration, interpretation and visualization is bespoke.
Phase 2a – Biobank federation
- federate global biobank data on up to 50 million individuals from diverse ancestries
- apply PRS from Phase 1 across biobanks to probe phenotype definitions and trans-ethnic heritability
- perform Phenome-wide Association Study (PheWAS) for all trait-associated alleles from Phase 1
- for associated alleles within “solved genes”, perform PheWAS to quantitatively model pleiotropic consequences for dose-response curves
- create searchable database, including visualization tools for dose-response portal, for all results
- implement GA4GH data sharing methods and create data sharing infrastructure for all future biobank studies
- create modular, open-source, automated pipeline for all components above to incentive future biobanks to federate data
Rationale: As with GWAS data, current biobanks are silo’d, which limits ability to conduct cross-biobank analyses; most clinical traits are not represented in existing cohort-based genetic studies; no standardize data sharing format.
Phase 2b – Variant to function mapping
- generate single-cell eQTL data in pathogenic cell types in >500 individuals; compare magnitude of eQTLs across cell types; incorporate data into dose-response portal
- generate exome sequencing and perform rare-variant association tests (RVAS) in >100,000 cases (selected from existing case-control cohorts and federated biobanks) in up to 10 selected diseases and/or quantitative traits; define genetic architecture using common variant association studies (CVAS) and RVAS; expand list of “solved genes” based on RVAS; perform PheWAS for disease-associated rare variants.
- perform high-throughput CRISPR perturbations in relevant cellular systems to experimentally validate causal variants from “solved genes”; reverse-engineer network wiring and define critical nodes within regulatory pathways; estimate magnitude of biological effect of causal variants; validate cellular readouts for future phenotypic screen
- conduct a “human knock-out project” via whole genome sequencing in >20,000 individuals from consanguineous populations and annotation all putative loss-of-function mutations
- utilize all data above to refine dose-response portal
- nominate genetically-validated cellular systems and perform phenotypic screen with all approved drugs; map all “solved genes” to regulatory nodes; estimate fraction of solved genes that are perturbed by approved drugs; catalogue new therapeutic modalities that would be required to target novel nodes.
Rationale: experimental validation is required to confirm pathogenic cell types, and to estimate quantitatively magnitude of biological effect of causal variants; need to understand complete genetic architecture (common to rare variants) of a selected set of diseases; need to understand phenotypic consequences of human knockouts; there is no map of genetic nodes and approved therapies.
So what are we waiting for? Let’s get started!